Toolbox: Putting corona on ice
The 2011 film Contagion became a popular streaming choice in the early days of the COVID-19 pandemic. It was lauded as a realistic depiction of the deadly fallout from a new virus that spreads through respiratory droplets. But David Kern, a postdoc in Steve Brohawn’s lab in the molecular and cell biology department at UC Berkeley, keeps thinking about the plot point that seemed least plausible when the movie first hit theaters. “They resolved the structure of a viral [protein] insanely quickly, and I thought: there’s no way they would be able to know this so soon,” he says. Nine years after Contagion’s release, that part seems less far-fetched as his research group shares the results of their rapid characterization of a small protein called 3a, which plays a crucial role in the lifecycle of SARS-CoV-2, the virus responsible for COVID-19.
Kern and Chris Hoel, a graduate student who was also part of the study, are amazed by the “resolution revolution” in cryogenic electron microscopy (cryo-EM) that has transformed their field over the past decade. As structural biologists, they determine a protein’s 3D structure, which, in turn, elucidates its function. Before high-powered computers and high-resolution cameras, scientists had to dissolve many copies of a given protein in water and coax them into forming a stable crystal to get a good picture of the molecule. The problem is that some proteins can take months to crystallize, while others, particularly those like 3a that dissolve in fats instead of water, never do. Cryo-EM, on the other hand, generates a 3D protein structure by stitching together 2D snapshots of a single molecule after it has been flash frozen, circumventing the challenge of crystallization.
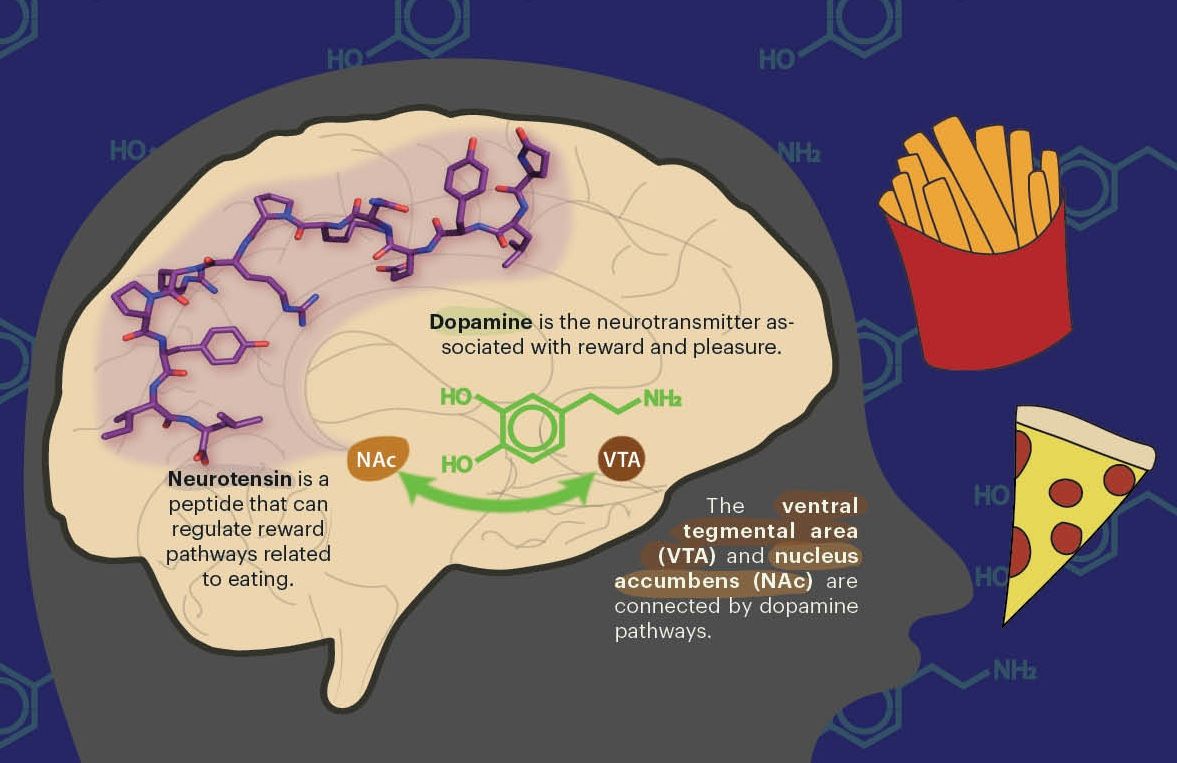
The Brohawn lab is one of several research groups around the world that have focused the power of cryo-EM on the proteins of SARS-CoV-2. Their goal is to identify specific viral proteins that are central to a virus’ lifecycle, which can be disrupted with antiviral drugs. The genetic sequence of the 3a protein identified it as a potential viroporin, a viral protein that creates a channel through fatty cell membranes and allows ions—charged particles such as calcium— to pass through. The ionic concentrations within a human cell are not homogenous because the cell actively moves ions between distinct subcellular regions separated by membranes. This heterogeneity is essential for healthy functioning. A cell that loses control of its ionic concentrations may die and burst open, releasing its contents into the surrounding tissue. By making membranes permeable to ions, viroporins can facilitate the release of a virus’ progeny, which can then infect neighboring cells.
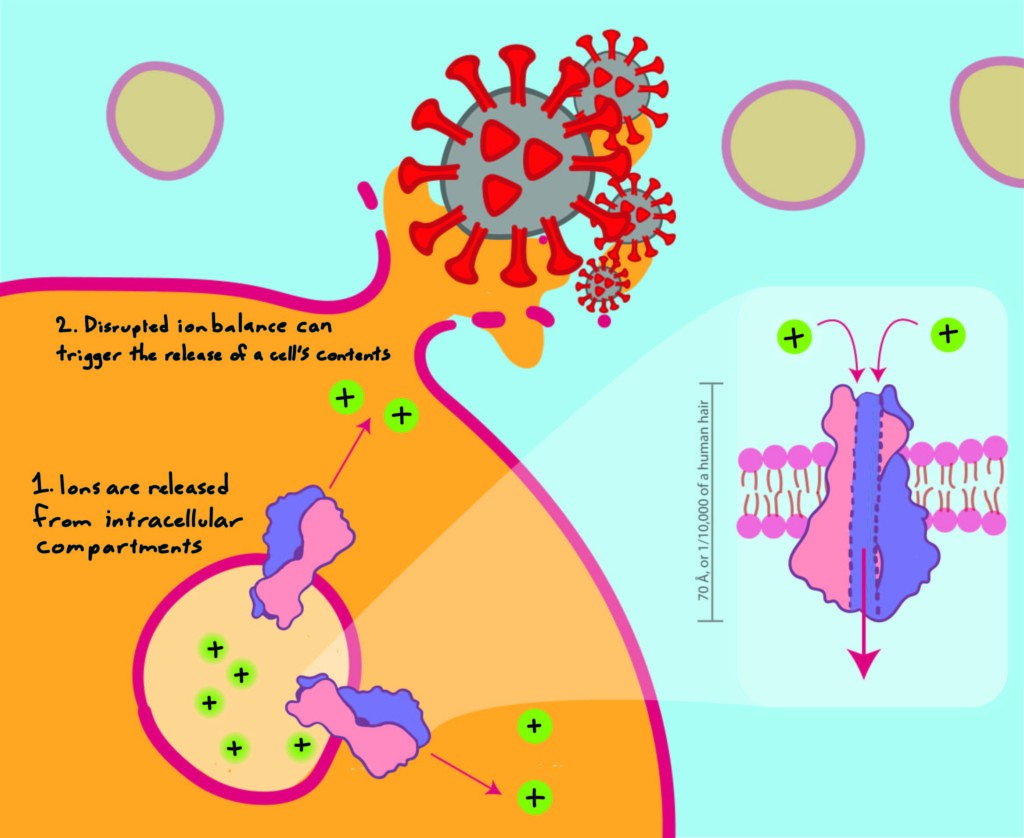 The 3A protein makes the host cell permeable to ions, which facilitates the release of progeny virus.
The 3A protein makes the host cell permeable to ions, which facilitates the release of progeny virus.
To study 3a, Kern and his team inserted the viral gene into cultured cells, which produced the protein for them. The team then isolated the 3a protein from the cells and tested its permeability to ions, ultimately confirming that it was a viroporin that let through positively charged ions like calcium. Finally, they fired up the electron microscope and resolved its structure. Just a decade ago, biologists may not have even tried to deter- mine the structure of a protein like 3a, but the whole process, from inserting the gene into the cells to producing a 3D model, took about a month to complete. Now the Brohawn lab is working together with its partners in other UC Berkeley labs to identify portions of the 3a protein’s structure that can be disrupted by existing antiviral drugs. “Because of the very recent resolution revolution in cryo-EM, these targets are more accessible, whereas during the original SARS [outbreak in 2002], [the SARS-CoV-1 virus] just would have been that much more challenging for people to try to study,” says Hoel. It can be hard to appreciate as the pandemic drags on, but the pace of scientific discovery keeps accelerating.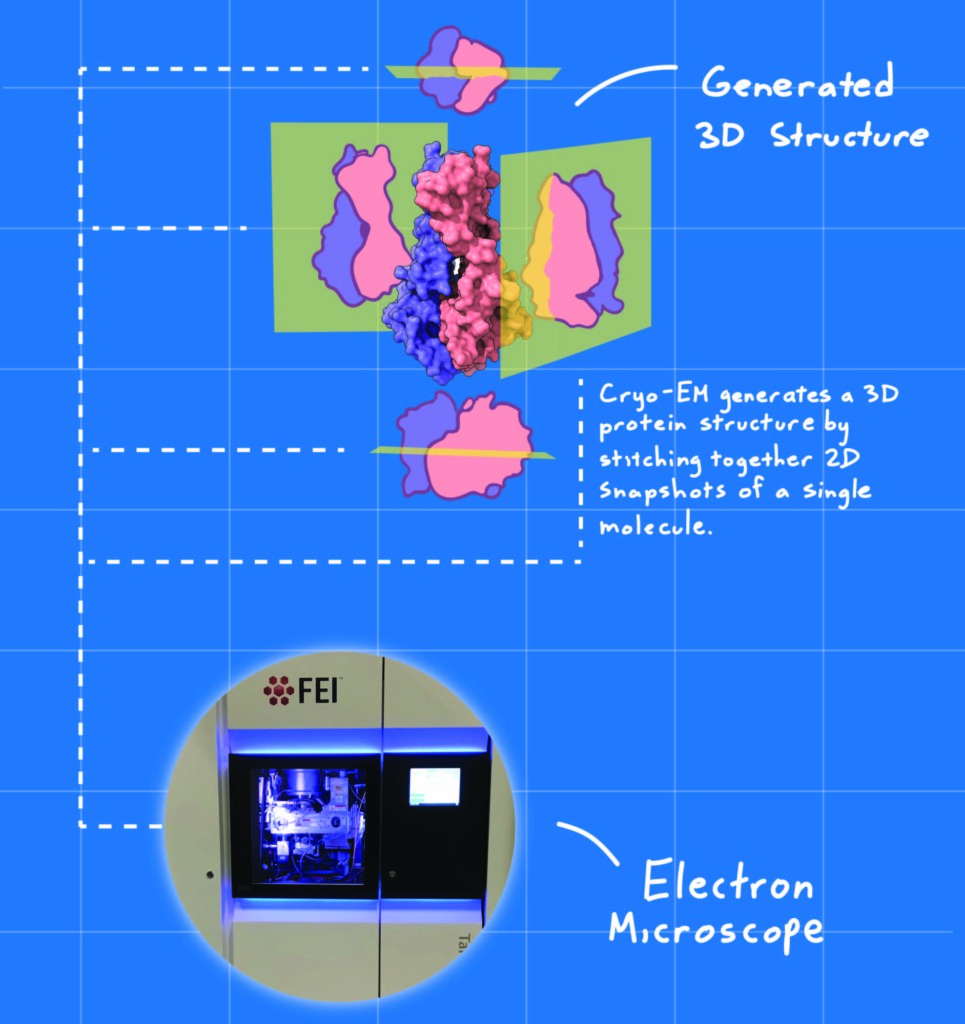
[divider type="thin"]
Andrew Saintsing is a graduate student in integrative biology
This article is part of the Fall 2020 issue.